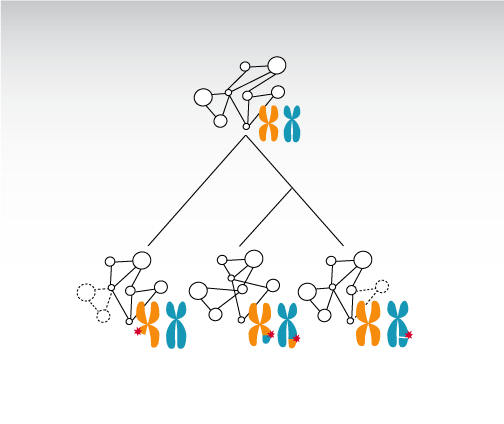
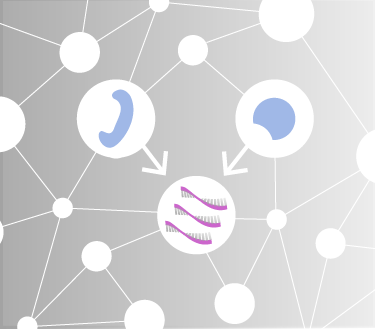
Living systems are made up of many different types of molecular entities, and accordingly there are different types of networks. One of the most important types of networks exists at the layer of “gene regulation”, which controls what set of genes get expressed at any point in time and space. Our work has focussed primarily on understanding the transcriptional layer of gene regulation.
Research in our group has progressed in three main directions and we are pursuing a number of questions under each major direction:
Gene regulatory network structure and function

3D genome organization
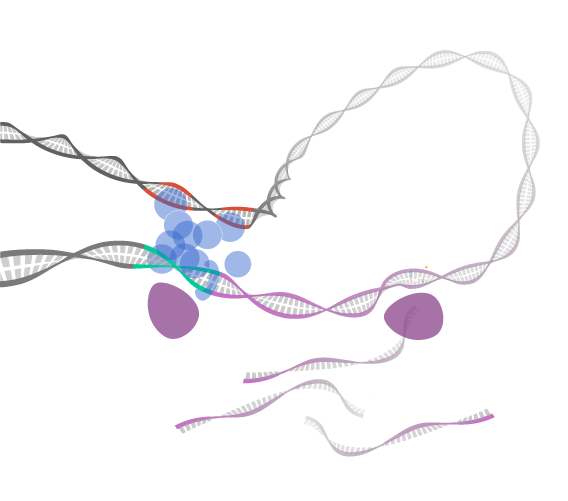
Evolution of gene regulation and regulatory networks